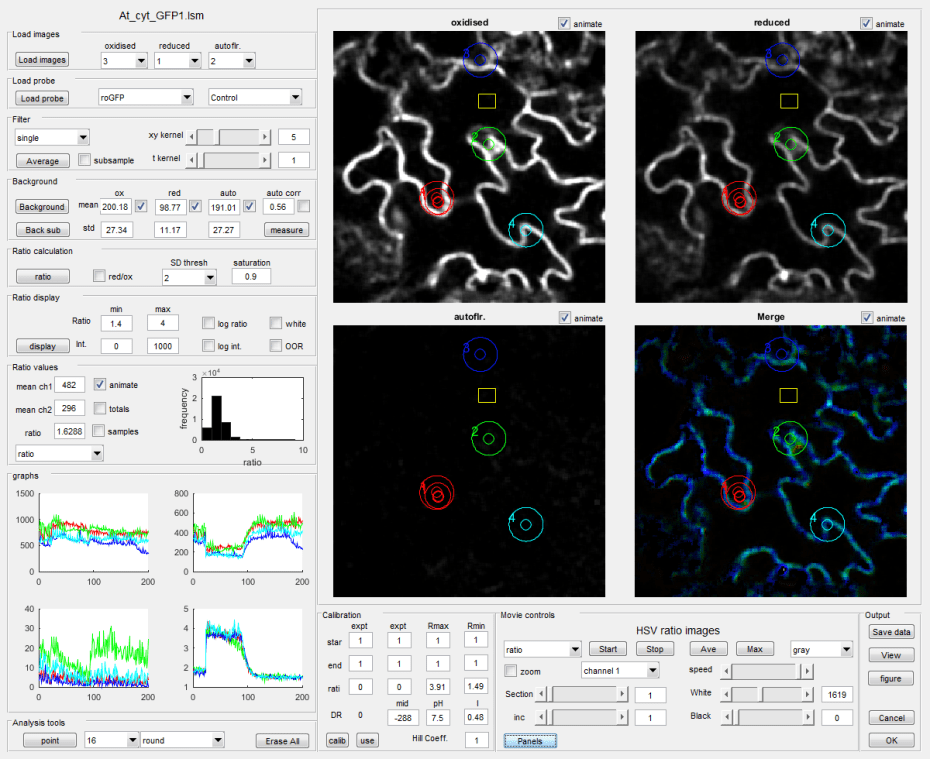
Basic Ratio Analysis
The basic ratio analysis program is designed to analyse single (x,y) images, 3-D (x,y,t) time series or 3-D (x,y,z) z-stacks. The first processing step usually involves improving the signal-to-noise ratio (S/N) by spatial or temporal averaging. Whilst there are a range of noise-reduction algorithms available, simple averaging using 2-D or 3-D kernels of different sizes provides a balance between S/N and spatio-temporal resolution. Averaging can be usefully combined with sub-sampling to reduce image sizes and increase processing speed, as spatial resolution is already compromised by averaging. As averaging is likely to yield non-integer values, this step also requires conversion from integer to floating-point format.
Outline of approach
To measure intensity values correctly it is important that the images are collected with a non-zero background, so that the statistical properties of the background can be estimated accurately and subtracted from each channel. In addition, auto-fluorescence bleed-through into one of the measurement channels is a common problem, particularly with plant specimens and blue/violet excitation. As auto-fluorescence has structure in the image, it cannot simply be subtracted as a single value like the background. However, it is possible to estimate the auto-fluorescence from an emission wavelength range that does not have any signal from the probe. For example, with roGFP, this is possible with excitation at 405 nm and emission at 435-485 nm. As the auto-fluorescence spectrum tends to be quite broad, a scaled version of the auto-fluorescence image can be subtracted from the probe image to correct for the auto-fluorescence bleed-through, provided the auto-fluorescence spectrum does not alter with time or treatment.
It is also useful to mask pixels with intensity values close to background or near saturation from the ratio image, or pixels that are drawn from regions with high local coefficient-of-variation (CV) that might correlate with edges of structures or compartments that are not well resolved, as these give a noisy and mis-leading impression of the real ratio value. Without masking, edge effects from ratioing low signal intensities can be substantial and can visually dominate the ratio image in plant and fungal specimens.
For pseudo-colour display, the masked ratio is coded by hue on a spectral colour scale ranging from blue (fully reduced) to red (fully oxidised), with the limits set by an in situ calibration or extrapolated from an in vitro calibration. The most useful initial mapping is to HSV (Hue, Saturation and Value or intensity) colour space, where the ratio is coded as Hue and the average intensity from both channels gives the Value. This is converted back to RGB colour space for display, and effectively gives bright colours for regions with good signals that fade to background for regions of low signal. If the data is a time-series, it can be viewed as a movie or montage. If the original data is a z-stack, visualisation benefits from calculating rocking or tilting projections to aid 3-D interpretation via motion parallax. Conventional rotation algorithms use maximum brightness projections of RGB images. However, this is not appropriate for colour-coded ratio images. Instead, projections in HSV colour-space are used to record the (x,y,z) position of each voxel in maximum projection of the V channel alone. The corresponding voxel from the H and S channels is then used to reconstruct the rotated pseudo-colour image.
Whilst ratio images are a useful tool to help visualise spatial variation in signal responses, the signal-to-noise (S/N) ratio is extremely low for single pixels, even after spatial and temporal smoothing. Quantitation is better achieved by manually selecting larger regions-of-interest (ROIs), which have some morphological or physiological significance. Each ROI encompasses many pixels with corresponding improvement in signal-to-noise ratio (S/N) at the expense of spatial resolution, that can be easily presented and interpreted in graphical form.
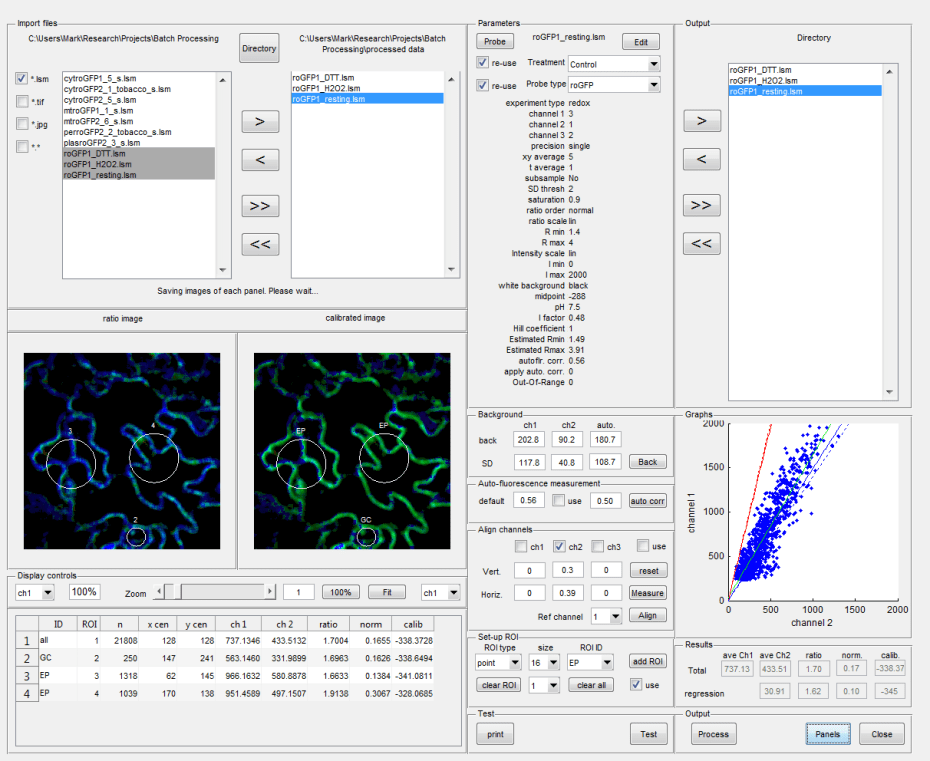
Batch Ratio analysis
The basic ratio program also has an option, to batch process a complete set of single (x,y) images present in one folder which is useful to compare the steady-state redox potential in multiple samples and treatments, measured over the whole image or from defined ROIs, rather than a continuous evaluation of changes in redox potential in one specimen.